Functional imaging of signaling networks
Understanding cellular and molecular mechanisms of signaling specificity
Research
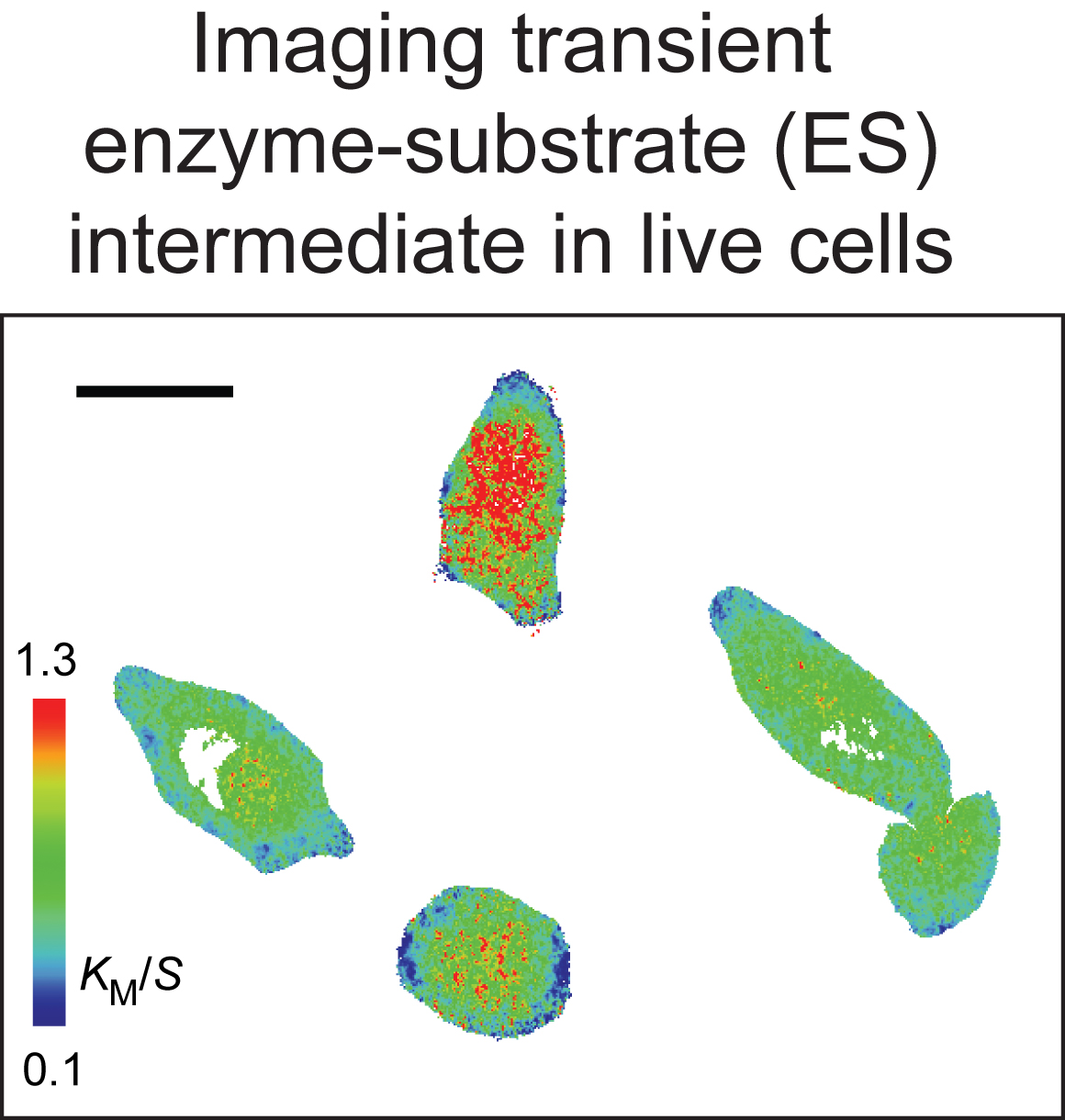
Adaptive character of responses to signals from the environment is a fundamental property of all living organisms. At the cellular level, it is brought about by a highly integrated process of transmembrane and intracellular signal transduction. Although many signaling molecules have been identified, how exactly the dynamics of their interactions ensure the ultimate specificity and adaptive character of cellular responses remains poorly understood. How changes in flux of substrates via an enzyme affects signalling specificity? How intracellular localization of a protein is coupled to its signaling role? What is the function of a given protein in network regulation?
To address these questions, we employ a combination of biochemical and advanced imaging techniques. We are developing tools to visualize the functional state of signaling molecules in live cells to determine how spatial distribution, regulation of specific activity and the corresponding changes in the flux of substrates via individual enzymes determine their signaling function. Furthermore, to examine the physiological relevance of individual enzymes in signaling, we will be developing new tools to selectively manipulate their localization and activity in live cells.
RESEARCH HIGHLIGHT:
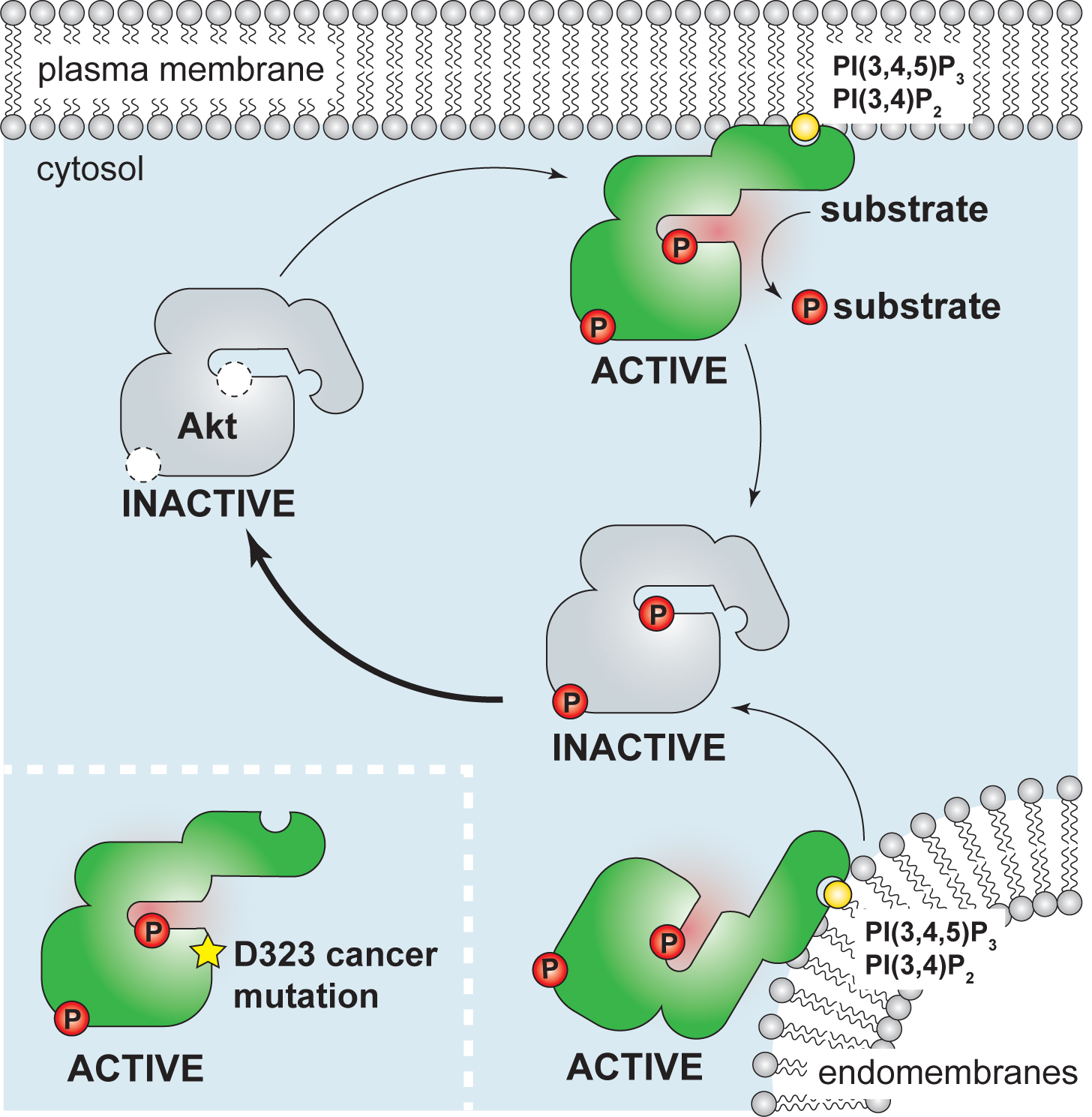
The new model of Akt activation in cells
In the recent paper in the journal Molecular Cell, Ebner et al show that binding to the membrane lipid PI(3,4,5)P3 allosterically activates Akt kinase by relieving an intramolecular autoinhibitory mechanism. Ebner et al further demonstrate that in cells active Akt is exclusively associated with PI(3,4,5)P3-containing cellular membranes, ensuring that Akt activity remains proportional to stimuli upstream of PI(3,4,5)P3.
Full text PDF PDF + Supplemental Materials
The constructs used in the publication are now available via AddGene
The primary data related to the publication are available at FigShare
Development of reporters to examine signaling dynamics in live cells
With the goal of examining regulation of enzymes in the complex cellular environment, we continuously develop optical and biochemical reporters, suitable for monitoring and manipulating the localization and activity of enzymes in live cells. We actively employ fluorescence lifetime imaging microscopy (FLIM), fluorescence correlation- and cross-correlation spectroscopy (FCS/FCCS), chemical and optogenetic probes and computer simulations to gain insight into operation and regulation of enzymes in cells.
Cellular regulatory mechanisms of protein kinase Akt
Protein kinase B, also known as Akt is a central regulator of cellular metabolism, survival and proliferation. It integrates signals from cell surface receptors coupled to PI3 kinase and links them to downstream signalling pathways by phosphorylating multiple (>200) substrates at a conserved motif. When activity of Akt is not properly controlled, this could lead to uncontrolled cell division and enhanced survival, as observed in many human cancers.
Using a combination of imaging, biochemical reporters and in vitro studies with purified Akt, we characterize cellular and molecular mechanisms which restrain Akt activation and control its substrate specificity in cells.
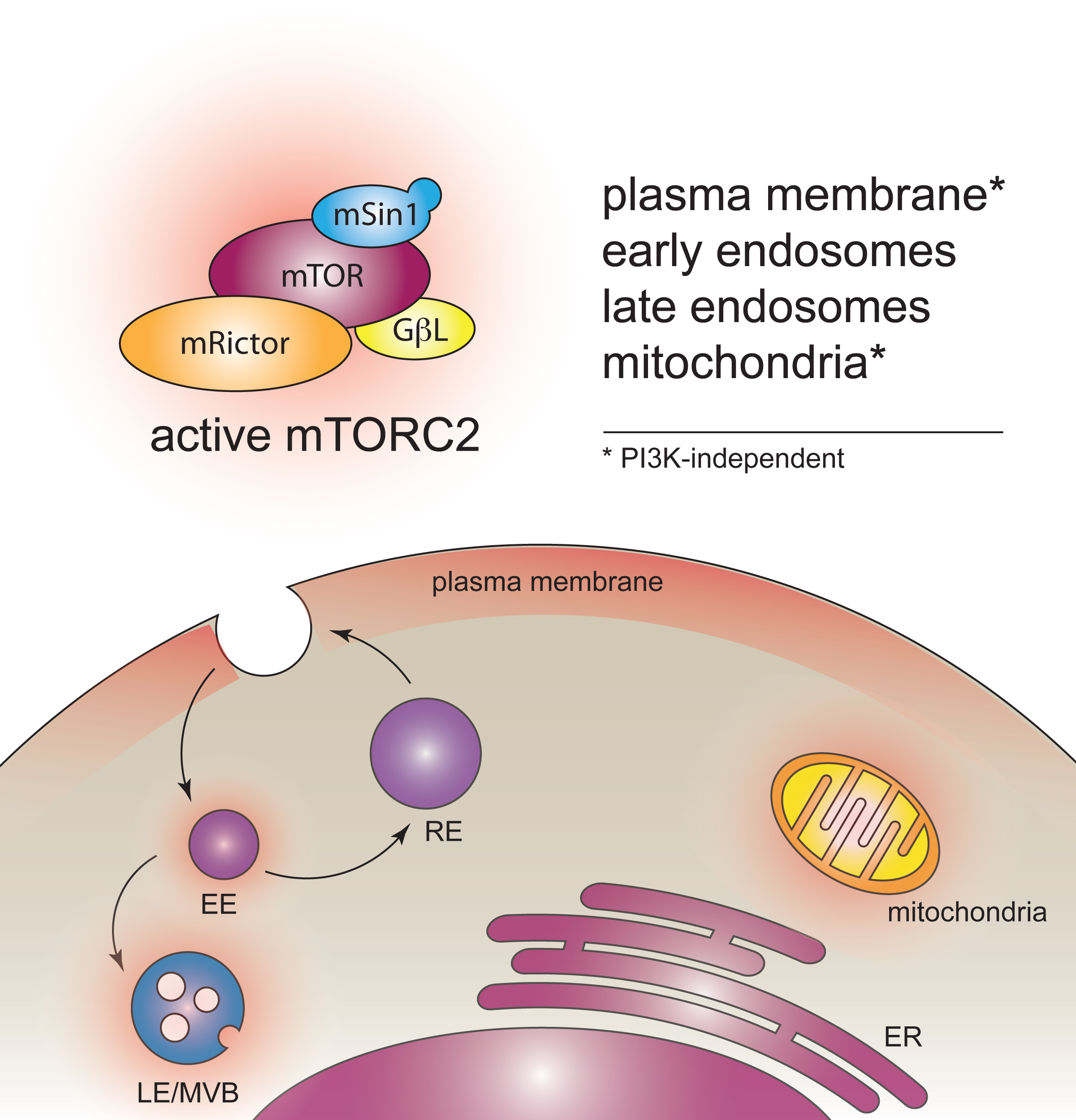
Intracellular localization and regulation of mTOR complex 2
Mechanistic target of rapamycin (mTOR) couples signalling from growth factor receptors with intracellular nutrient-sensing pathways, and thereby is a critical regulator of cellular anabolic functions, growth, survival and proliferation. In eukaryotic cells, mTOR exists in two distinct multisubunit complexes, mTORC1 and mTORC1, which have non-overlapping functions in cellular physiology. Specifically, mTORC2 is implicated in the control of cell proliferation, survival, cytoskeletal rearrangements and, indirectly, in regulation of mTORC1-controlled signalling.
Unlike mTORC1, little is known about localization and regulation of mTORC2. We use combination of superresolution imaging and specifically designed biochemical reporters to examine localization and regulation of mTORC2 activity inside cells.
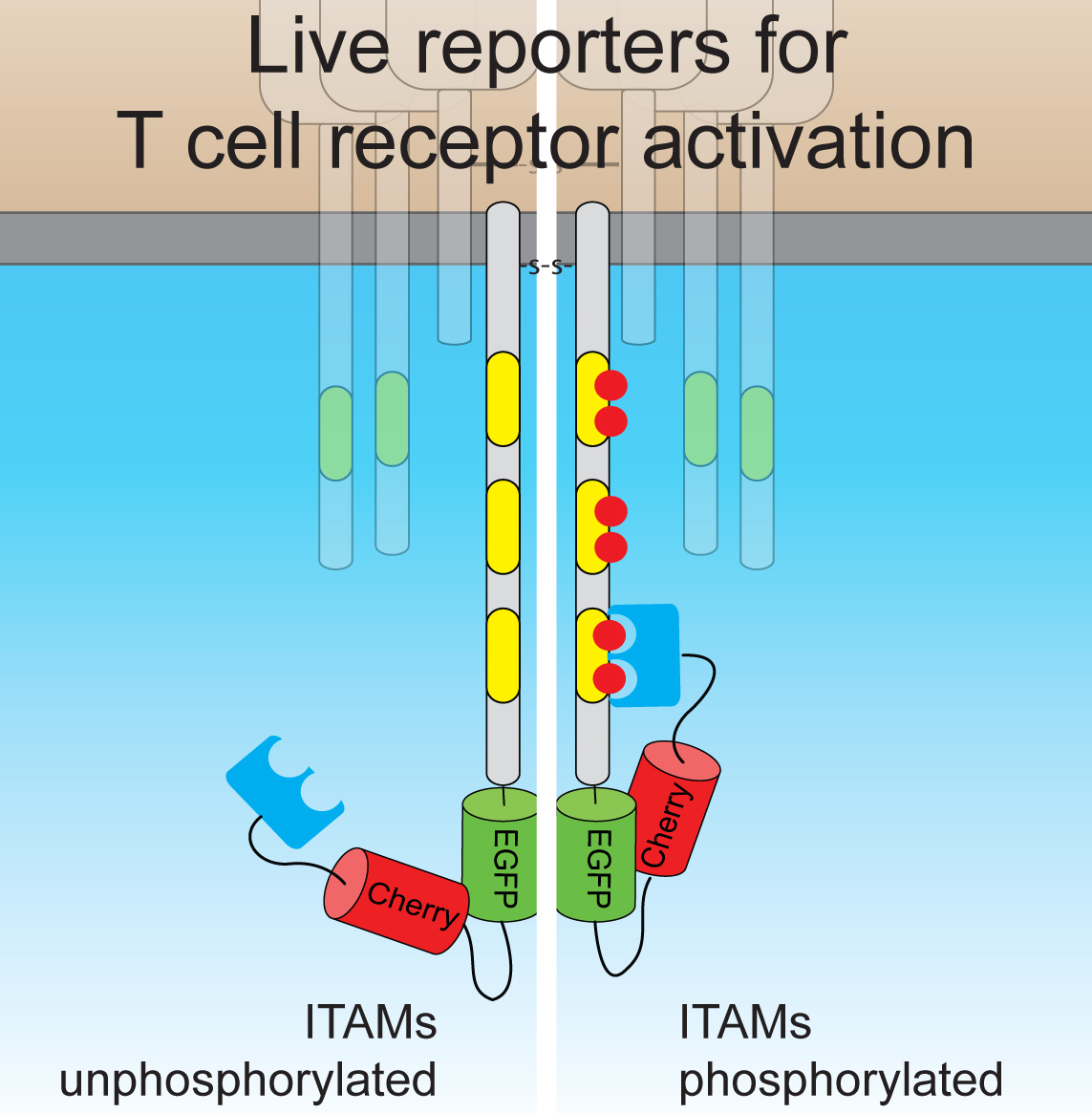
Dynamics of T cell receptor signaling
T cell receptor complex (TcR/CD3) mediates physiological responses of lymphocytes to antigenic stimulation. Antigen binding induces phosphorylation of the cytoplasmic portion of the TcR/CD3 and subsequent recruitment and activation of the cytosolic kinase ZAP-70. Activated ZAP-70 in turn phosphorylates scaffold proteins LAT and SLP-76 resulting in recruitment of downstream effectors, which link TcR/CD3 activation with cytokine secretion, cytoskeletal rearrangement, mitogenic signalling and gene expression.
Similar to other transmembrane receptors, TcR/CD3 is constitutively internalised and recycled to the plasma membrane in T cells, although the exact role of its constant trafficking inside cells is unclear. We use imaging and biochemical approaches to investigate TcR/CD3 signalling from subcellular compartments and examine its role in T cell physiology.
NEW OPEN ACCESS CRITICAL REVIEW:
Control of Akt activity and substrate phosphorylation
In the recent review in IUBMB Life, Ivan examines how the cell-autonomous and intrinsic allosteric mechanisms cooperate to ensure localized, context-specific signaling by Akt
NEWS: ASCB | EMBO 2017 Special Interest Subgroup
Brendan Manning (T.H. Chan School of Public Health, Harvard U) and Ivan Yudushkin are co-organizing a Special Interest Subgroup Spatial and temporal control of cell signalingat the upcoming ASCB | EMBO 2017 Meeting in Philadelphia.
Coordinated assembly, localization and regulation of enzymatic complexes shape cellular signaling responses to the environmental cues. To ensure the fidelity of these reactions, the cells need to precisely position the enzymes and their substrates at specific compartments and provide mechanisms to trigger and quench the enzymatic reactions in time. The details of the biochemical architecture and coupling between subcellular localization and regulation of signaling enzymes are not well understood.
The speakers of this Special Interest Subgroup include Wade HARPER (Harvard), Peter DEVREOTES (John Hopkins), Brenda ANDREWS (U Toronto), Jin ZHANG (UCSD), Anne-Claude GINGRAS (Lunenfeld), Roshanak Irannejad (UCSF), Oliver ROCKS (MDC Berlin) and Ivan himself. The speakers will present new methods for fast, high-resolution mapping, quantitation and modeling of cellular biochemical activities and discuss how dynamic architecture of enzymatic activities defines the adaptive character and fidelity of cellular responses.
The program of the Subgroup is available online. REGISTER EARLY to attend this exciting session on Saturday, December 2 2017, 8:30-12:30, Room 113A
This subgroup is kindly supported by the Journal of Cell Biology and Science Signaling

Chiara TRILLING, M.Sc student

Lukáš GULLA, M.Sc student

Michael EBNER, alumnis Ph.D. student

Benjamin SINKOVICS, alumnis M.Sc. student

Magdalena SZCZYGIEŁ, alumna summer student

Daniela WOLFSCHOON RIBEIRO, alumna rotation student

Ivan YUDUSHKIN, group leader
Selected publications
Far far away, behind the word mountains, far from the countries Vokalia and Consonantia, there live the blind texts.




Easily find and use practical tips for your current writing tasks
Modules and extensive cross-reference help you find the relevant information fast and start writing straightaway.
Better design
Far far away, behind the word mountains, far from the countries Vokalia and Consonantia, there live the blind texts. Separated they live in Bookmarksgrove right at the coast of the Semantics, a large language ocean.
Understand the principles
Understand the functions of each section in the manuscript, presentation or figure and learn how to pose questions, present evidence and formulate conclusions.
Plan ahead and track progress
Formulate achievable goals, outline the actionable plan and track progress. Learn simple tricks to avoid procrastination, writer's block and perfectionism.
Follow practical tips
Use provided questions, cues and checklists to explore your topic in full. Get inspired and enrich your vocabulary through examples.
Use expert tools
Learn about professional word processors, illustration, bibliography and typesetting software to help you along the way. Check the available resources in print and online.

Resources
Primary data
References to primary data for published articles on FigShare:
JCB LocaTOR2 paper
Molecular Cell Akt paper
Protocols
Protocols and references
Materials
References to Yudushkin lab plasmids deposited at AddGene:
LocaTOR2 JCB paper
Mol Cell paper constructs
TcR reporters PNAS paper
Contact
Department of Structural and Computational Biology
Max F. Perutz Laboratories (MFPL)
University of Vienna
Vienna Biocenter, Campus Vienna Biocenter 5, Rm. 1.612
1030 Vienna AUSTRIA
Plans built for every one
Far far away, behind the word mountains, far from the countries Vokalia and Consonantia, there live the blind texts.